FAQs
What your patients need to know

General queries
Information on which clinical specialties can order a specific genomic test is detailed in the National Genomic Test Directory.
For Rare and Inherited Diseases, this information is found in the Eligibility Criteria document. If you are unsure whether you are able to order a test please contact us: gos-tr.norththamesgenomics@nhs.net.
For rare and inherited disease tests, please check the Eligibility Criteria for the Genomic Test Directory.
For cancer tests, please check the eligibility criteria is listed in the National Genomic Test Directory for Cancer.
Referrals must meet these criteria and will be triaged at the Genomic Laboratory Hub.
For Rare and Inherited Diseases, information for sending a sample to the GLH lab based at Great Ormond Street can be found here.
For Cancer, please see The Royal Marsden’s Clinical Genomics Service Specification.
No. Tests listed in the National Genomic Test Directories are funded by NHS England.
Reporting times vary depending on the test type and referral type. The national guideline turnaround times for different types of test are listed in the table below. Please note that some tests may not meet these turnaround times currently. If you have requested a test and have not received the results by the turnaround times, please contact the laboratory for current turnaround timings gos-tr.norththamesgenomics@nhs.net.
Please check the turnaround times for the test you have ordered. If you have requested a test and have not received the results by the turnaround times, please contact the laboratory for current turnaround times: gos-tr.norththamesgenomics@nhs.net.
There may be different reasons for an inappropriate referral. Usually it is due to either an inappropriate sample (where the wrong sample has been received) or due to an issue with eligibility (either insufficient clinical details or a request from a specialist other than those permitted by the Test Directory). See our referral forms for sample types for different tests, and see the Test Directory Eligibility Criteria for further information about eligibility.
We provide testing for a diverse population of around 10 million people, across 29 NHS Trusts;
– Barking, Havering and Redbridge University Hospitals NHS Trust
– Barnet, Enfield and Haringey Mental Health NHS Trust
– Barts Health NHS Trust
– Camden and Islington NHS Foundation Trust
– Central and North West London NHS Foundation Trust
– Central London Community Healthcare NHS Trust
– Chelsea and Westminster Hospital NHS Foundation Trust
– East London NHS Foundation Trust
– Essex Partnership University NHS Foundation Trust
– Great Ormond Street Hospital for Children NHS Foundation Trust
– Hertfordshire Community NHS Trust
– Hertfordshire Partnership University NHS Foundation Trust
– Homerton University Hospital NHS Foundation Trust
– Imperial College Healthcare NHS Trust
– London North West Healthcare NHS Trust
– Mid and South Essex Hospital Services NHS Trust
– Moorfields Eye Hospital NHS Foundation Trust
– North East London NHS Foundation Trust
– North Middlesex University Hospital NHS Trust
– Royal Free London NHS Foundation Trust
– Royal National Orthopaedic Hospital NHS Trust
– Tavistock and Portman NHS Foundation Trust
– The Hillingdon Hospitals NHS Foundation Trust
– The Princess Alexandra Hospital NHS Trust
– The Royal Marsden NHS Foundation Trust
– The Whittington Hospital NHS Trust
– University College London Hospitals NHS Foundation Trust
– West Hertfordshire Hospitals NHS Trust
– West London Mental Health NHS Trust
We also provide cancer testing for;
– Kingston Hospital NHS Foundation Trust
– Croydon Health Services NHS Trust
– Epsom and St Helier University Hospitals NHS Trust
The NHS GMS has established a network of 7 regional genomic laboratory hubs, each responsible for coordinating and delivering genomic testing for a particular part of the country.
We work cooperatively with the national network of GLHs to deliver all tests outlined in the National Genomic Test Directory and bring equitable access to testing across the country.
– NHS Central and South GLH
– NHS East GLH
– NHS North West GLH
– NHS South East GLH
– NHS South West GLH
– NHS North East and Yorkshire GLH
Please visit our Education and Training section.
National Genomic Test Directory
Please visit our National Genomic Test Directory (NGTD) page.
The Test Directory can be filtered to make it easier to find tests relating to your specialty. You may find that some tests that you previously ordered now have a different name in the Test Directory. If you are having difficulty finding the relevant test please contact the laboratory: gos-tr.norththamesgenomics@nhs.net.
The National Genomic Test Directory (NGTD) outlines all genetic tests funded by NHS England, and is developed by expert groups including clinicians, scientists, health economists, policy experts, public representatives and patient organisations.
Any genomic tests not included are out of scope of the GLH. The NGTD will be updated annually as evidence arises to suggest the inclusion of new genes. Information about the process for updating the NGTD can be found on the NHS England website.
NHS England have also published a National Genomic Test Directory Frequently Asked Questions (FAQ) document.
The NGTD is updated annually as evidence arises to suggest the inclusion of new genes. Information about the process for updating the NGTD can be found on the NHS England website.
Applications to request new tests are usually open between June-August each year, and the GLH is happy to support clinicians with their submissions.
Rare and Inherited Conditions
There is a Test selection Tool NHS England | Public Genetic Test Directory (genomics.nhs.uk) if you enter the disorder in the search box, a list of suggested panels is generated.
By clicking on a suggested panel – under further info a list of genes that are included is generated.
For DNA-based tests you can request additional tests where we have DNA stored already. Please complete a new request form and forward it to genetics.labs@nhs.net.
Yes you can, as long as your primary clinical indication falls within the Phase 1 and Phase 2 indications listed in the table. Then, if your patient has other symptoms, too, you can add one of the panels which are listed as ‘GMS Rare Disease Virtual’ on panelapp e.g. Hearing loss panel.
Results are returned by Outreach portal or email. Results will be posted in rare circumstances.
Outreach is a results portal used by the Rare & Inherited Disease Laboratory to issue reports to referring departments. Once a report is verified it is sent automatically to the Outreach portal, and the email address registered to the account will receive a notification that a report is available. Using Outreach means you do not need to wait for reports to be emailed to you by the lab’s administrative team.
If you would like to begin using Outreach, please contact gos-tr.norththamesgenomics@nhs.net to request a form.
Please note, you will need to ensure that the email address you provide is a shared/departmental account, NOT a personal email address. This is so that reports are available to other colleagues in the event that you are away from work. Please also note that as a standard, we do not also email reports that are configured to be delivered via Outreach.
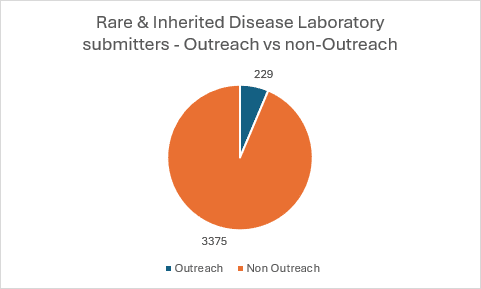
Data around current Outreach utilisation is below:
Whole Genome Sequencing
In general, it is recommended to send a trio (affected patient plus parents) for WGS. This makes it easier to analyse the variant data and more likely to reach a diagnosis than with a singleton referral.
However, there are many circumstances where parents may not be available and the case can be referred as a singleton or a duo.
Yes. Patients and family members can make a Subject Access Request (SAR) to have their raw genomic data. Clinicians should inform patients that this data would need to be interpreted by a genomics expert and is not presented as a clinical report outlining all genomic changes. The clinician should contact the WGS service desk at gos-tr.wgsnorththamesglh@nhs.net if their patient wishes to make a SAR.
For Rare and inherited conditions, a Germline Sample is required: peripheral blood (2-5ml EDTA), stored DNA, cultured fibroblasts.
For cancer a Germline Sample is required: peripheral blood (2-5ml EDTA), stored DNA, cultured fibroblasts, for haematological malignancies a skin biopsy is required. A tumour sample is also required:
- For Solid tumours: Fresh frozen tissue (at least 1 core biopsy), tumour cells must account for at least 30% of the nucleated cells present
- For haematological malignancies: Bone marrow aspirate, peripheral blood
Proband and both parents is the preferred family structure as adding affected siblings can complicate the analysis. If a pertinent result is found through WGS, targeted testing may be offered to affected siblings, if appropriate.
Clinicians will receive a report once analysis is complete. You may be asked to provide further information or attend an MDT / GTAB before the report is issued.
If the policy was taken out before the patient received their test result, they are not obliged to update their provider on the test and its findings for as long as the cover is in place and remains unaltered. Further information on the patients current and future responsibility towards insurance providers is provided by Genetic Alliance UK and the Association of British Insurers. Code on Genetic Testing and Insurance ABI.
To find out more about which insurance companies are registered with the ABI and subject to the above terms please visit: ABI members.
Analysis for WGS is panel based so please ensure that the correct clinical indication is entered in the Test Order Form so that the right panels are applied. For Rare Diseases, clinicians are able to add additional panels, if appropriate.
Reanalysis is available where there is clinical need and where reanalysis is likely to increase the chance of a diagnosis due to new gene associations, additional clinical information, improved bioinformatics.
Requests for WGS data reanalysis may be accepted as appropriate if the following criteria are met:
There must be a high expectation that reanalysis will yield a diagnosis. For example:
- The change to gene panel content or bioinformatic improvements introduced since the last analysis was performed, is significant so that the probability of achieving a diagnosis is sufficient enough to justify reanalysis. It is not expected that reanalysis will be significantly productive within at least 2 years of the last analysis.
AND
There must be a clinical trigger for the reanalysis. For example:
- There is a new significant change in the clinical presentation of the patient or family member (e.g. evolving phenotype) meaning application of additional GMS approved gene panel(s) would be relevant and updating the HPO terms provided will support more effective analysis in Exomiser;
- New treatment becomes available so reanalysis could influence the patient’s clinical management;
- The proband is deceased and the family initiate further investigation through the appropriate clinician, due to potential management implications for themselves and/or the wider family.
If you consider that re-analysis should be performed please contact the laboratory: gos-tr.wgsnorththamesglh@nhs.net
Patient’s DNA samples should not be sent for resequencing via the current GMS WGS pathway apart from in specific circumstances.
Outlined below are possible exceptions for which it is acceptable to submit patients and their family members for WGS through the GMS pathway:
- Patient submitted to 100,000 Genomes Project as a singleton but it is now possible to test as a Trio (for clinical indications whereby trio analysis is the optimal family structure) and the primary analysis detected lots of potential candidate variants which trio analysis will help to classify.
- There is immediate clinical urgency AND an expectation that a new test would yield a new diagnosis, e.g. The original analysis was done several years ago, the phenotype has substantially altered and a molecular diagnosis is likely to contribute to urgent clinical decisions. The patient can be tested through the relevant test from the GMS National Test Directory (including WGS if clinically eligible).
Yes.
National Genomic research library (NGRL)
The National Genomic Research Library (NGRL) is a national research database owned by Genomics England which holds the genomic, health and sample data of patients who have undergone whole genome sequencing (WGS). All rare disease and cancer patients who are consented for WGS have the opportunity to share their anonymised data with this library.
This anonymised data is vital for advancing research, helping us to improve our understanding of human disease and develop treatments that can benefit patients with many conditions. More information about the NGRL can be found on the Genomics England website (https://www.genomicseducation.hee.nhs.uk/supporting-the-nhs-genomic-medicine-service/national-genomic-research-library-information-for-clinicians/)
The NGRL allows approved access to the pseudonymised health data, genomic data and samples of patients and family members who have had WGS and consented to their data being accessed to the library.
While patients can be identified if necessary (e.g. If a diagnosis is found or there is potentially relevant research), NGRL governance specifies that researchers will not be able to identify individuals.
You can apply to become a member of the academic research community (GECIP) by following this link https://www.genomicsengland.co.uk/research/academic/join-gecip